-
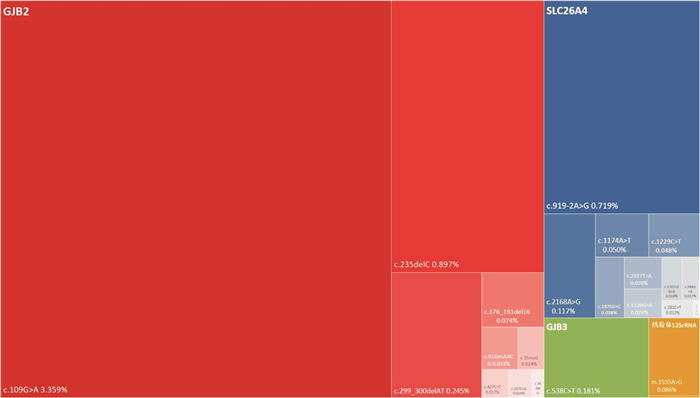
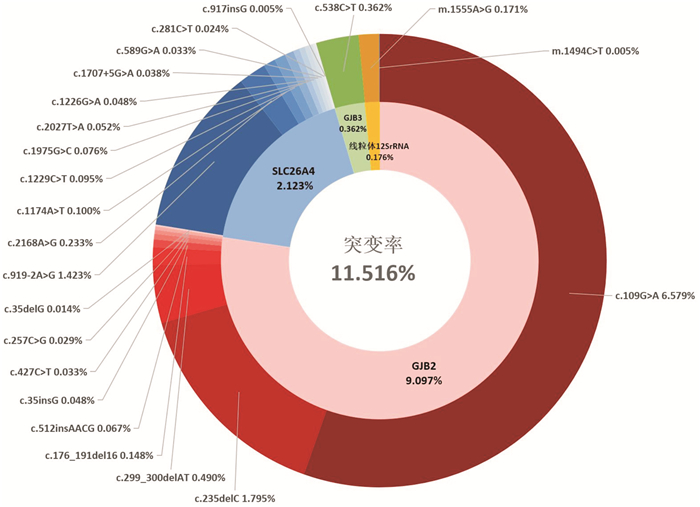
摘要: 目的 分析北京市23项新生儿耳聋基因筛查的突变频谱,为遗传咨询及临床诊疗提供依据。方法 研究对象为2022年12月-2023年6月在首都医科大学附属北京同仁医院接受23项耳聋基因筛查的新生儿21 006例。23项耳聋基因筛查包括4个基因23个位点: GJB2 基因(c.35delG、c.176_191del16、c.235delC、c.299_300delAT、c.109G>A、c.257C>G、c.512insAACG、c.427C>T、c.35insG)、 SLC26A4 基因(c.919-2A>G、c.2168A>G、c.1174A>T、c.1226G>A、c.1229C>T、c.1975G>C、c.2027T>A、c.589G>A、c.1707+5G>A、c.917insG、c.281C>T)、线粒体 12SrRNA 基因(m.1555A>G、m.1494C>T)和 GJB3 基因(c.538C>T)。分析各基因位点的突变率及等位基因突变频率。结果 21 006例中,耳聋基因筛查未通过率11.516%(2 419/21 006)。4个基因中 GJB2 基因突变率最高,为9.097%(1 911/21 006),其次分别为 SLC26A4 基因2.123%(446/21 006)、 GJB3 基因0.362%(76/21 006)及线粒体 12SrRNA 基因0.176%(37/21 006)。 GJB2 基因中,c.109G>A和c.235delC突变率最高,分别为6.579%(1 382/21 006)和1.795%(377/21 006)。 SLC26A4 基因中,c.919-2A>G和c.2168A>G突变率最高,分别为1.423%(299/21 006)和0.233%(49/21 006)。等位基因突变频率, GJB2 基因c.109G>A最高,为3.359%(1 411/42 012),其次为 GJB2 基因c.235delC,0.897%(377/42 012)及 SLC26A4 基因c.919-2A>G,0.719%(302/42 012)。结论 北京市23项新生儿耳聋基因筛查提示, GJB2 基因c.109G>A突变率和等位基因突变频率最高,值得临床重视。本研究丰富了23项新生儿耳聋基因筛查突变频谱的流行病学资料,可为临床提供依据。Abstract: Objective To analyze the mutation spectrum of 23-site chip newborn deafness genetic screening in Beijing, and to provide basis for genetic counseling and clinical diagnosis and treatment.Methods The study included 21 006 babies born in Beijing from December 2022 to June 2023. All subjects underwent newborn deafness genetic screening in Beijing Tongren Hospital, covering 23 variants in 4 genes, the GJB2 gene(c.35delG, c.176_191del16, c.235delC, c.299_300delAT, c.109G>A, c.257C>G, c.512insAACG, c.427C>T, c.35insG), SLC26A4 gene(c.919-2A>G, c.2168A>G, c.1174A>T, c.1226G>A, c.1229C>T, c.1975G>C, c.2027T>A, c.589G>A, c.1707+5G>A, c.917insG, c.281C>T), Mt12SrRNA(m.1555A>G, m.1494C>T) and GJB3 gene(c.538C>T). The mutation detection rate and allele frequency were analyzed.Results The overall mutation detection rate was 11.516%(2 419/21 006), with the GJB2 gene being the most frequently involved at 9.097%(1 911/21 006), followed by the SLC26A4 gene at 2.123%(446/21 006), the GJB3 gene at 0.362%(76/21 006) and Mt12SrRNA at 0.176%(37/21 006). Among the GJB2 genes, c.109G>A and c.235delC mutation detection rates were the highest, with 6.579%(1 382/21 006) and 1.795%(377/21 006), respectively. Of the SLC26A4 genes, c.919-2A>G and c.2168A>G had the highest mutation rates of 1.423%(299/21 006) and 0.233%(49/21 106), respectively. Regarding the allele frequency, GJB2 c.109G>A was the most common variant with an allele frequency of 3.359%(1 411/42 012), followed by the GJB2 c.235delC at 0.897%(377/42 012) and the SLC26A4 c.919-2A>G at 0.719%(302/42 012).Conclusion 23-site chip newborn deafness genetic screening in Beijing showed that GJB2 c.109G>A mutation detection rate and allele frequency were the highest. This study has enriched the epidemiological data of 23-site chip genetic screening mutation profiles for neonatal deafness, which can provide evidence for clinical practice.
-
Key words:
- newborn /
- deafness genes /
- mutation detection rate /
- allele frequency
-
-
表 1 23项耳聋基因筛查突变者的基因型分布
突变类型 基因型 突变例数 基因型占比(n=21 006) 纯合突变 c.109G>A/c.109G>A 29 0.138 c.919-2A>G/c.919-2A>G 3 0.014 复合杂合突变 c.235delC/c.109G>A 11 0.052 c.299_300delAT/c.109G>A 6 0.029 c.235delC/c.109G>A/c.538C>T 1 0.005 c.235delC/c.109G>A/c.919-2A>G 1 0.005 c.235delC/c.176_191del16 1 0.005 c.235delC/c.35delG 1 0.005 c.109G>A/c.35insG 1 0.005 c.919-2A>G/c.281C>T 1 0.005 线粒体12SrRNA突变 m.1555A>G均质 28 0.133 m.1555A>G异质 5 0.024 c.109G>A/m.1555A>G均质 2 0.010 c.109G>A/m.1494C>T均质 1 0.005 c.2168A>G/m.1555A>G均质 1 0.005 单杂合突变 c.109G>A杂合 1300 6.189 c.235delC杂合 351 1.671 c.919-2A>G杂合 273 1.300 c.299_300delAT杂合 95 0.452 c.538C>T杂合 68 0.324 c.2168A>G杂合 41 0.195 c.176_191del16杂合 30 0.143 c.1229C>T杂合 20 0.095 c.1174A>T杂合 19 0.090 c.1975G>C杂合 15 0.071 c.512insAACG杂合 13 0.062 c.35insG 9 0.043 c.1226G>A杂合 9 0.043 c.2027T>A杂合 8 0.038 c.427C>T杂合 7 0.033 c.589G>A杂合 6 0.029 c.257C>G杂合 6 0.029 c.1707+5G>A杂合 5 0.024 c.281C>T杂合 4 0.019 c.35delG 2 0.010 c.917insG杂合 1 0.005 双基因杂合突变 c.109G>A/c.919-2A>G 14 0.067 c.235delC/c.919-2A>G 6 0.029 c.109G>A/c.538C>T 5 0.024 c.235delC/c.2168A>G 4 0.019 c.109G>A/c.2168A>G 3 0.014 c.109G>A/c.1174A>T 2 0.010 c.109G>A/c.1707+5G>A 2 0.010 c.109G>A/c.589G>A 1 0.005 c.299_300delAT/c.538C>T 1 0.005 c.299_300delAT/c.919-2A>G 1 0.005 c.2027T>A/c.538C>T 1 0.005 c.109G>A/c.1226G>A 1 0.005 c.109G>A/c.1975G>C 1 0.005 c.109G>A/c.2027T>A 1 0.005 c.235delC/c.2027T>A 1 0.005 c.512insAACG/c.1707+5G>A 1 0.005 合计 2 419 11.516 表 2 4个基因23个位点的突变率
基因 位点 位点突变/例(%) 基因突变/例(%) GJB2 c.109G>A 1 382(6.579) 1 911(9.097) c.235delC 377(1.795) c.299_300delAT 103(0.490) c.176_191del16 31(0.148) c.512insAACG 14(0.067) c.35insG 10(0.048) c.427C>T 7(0.033) c.257C>G 6(0.029) c.35delG 3(0.014) SLC26A4 c.919-2A>G 299(1.423) 446(2.123) c.2168A>G 49(0.233) c.1174A>T 21(0.100) c.1229C>T 20(0.095) c.1975G>C 16(0.076) c.2027T>A 11(0.052) c.1226G>A 10(0.048) c.1707+5G>A 8(0.038) c.589G>A 7(0.033) c.281C>T 5(0.024) c.917insG 1(0.005) GJB3 c.538C>T 76(0.362) 76(0.362) 线粒体 m.1555A>G 36(0.171) 37(0.176) 12SrRNA m.1494C>T 1(0.005) 表 3 23个位点的等位基因突变频率
基因 位点 突变例数 杂合突变数 纯合突变数 等位基因突变数 等位基因突变频率/% GJB2 c.109G>A 1 382 1 353 29 1 411 3.359 c.235delC 377 377 0 377 0.897 c.299_300delAT 103 103 0 103 0.245 c.176_191del16 31 31 0 31 0.074 c.512insAACG 14 14 0 14 0.033 c.35insG 10 10 0 10 0.024 c.427C>T 7 7 0 7 0.017 c.257C>G 6 6 0 6 0.014 c.35delG 3 3 0 3 0.007 SLC26A4 c.919-2A>G 299 296 3 302 0.719 c.2168A>G 49 49 0 49 0.117 c.1174A>T 21 21 0 21 0.050 c.1229C>T 20 20 0 20 0.048 c.1975G>C 16 16 0 16 0.038 c.2027T>A 11 11 0 11 0.026 c.1226G>A 10 10 0 10 0.024 c.1707+5G>A 8 8 0 8 0.019 c.589G>A 7 7 0 7 0.017 c.281C>T 5 5 0 5 0.012 c.917insG 1 1 0 1 0.002 GJB3 c.538C>T 76 76 0 76 0.181 线粒体12SrRNA m.1555A>G 36 5 31 36 0.086 m.1494C>T 1 0 1 1 0.002 -
[1] Estivill X, Fortina P, Surrey S, et al. Connexin-26 mutations in sporadic and inherited sensorineural deafness[J]. Lancet, 1998, 351(9100): 394-398. doi: 10.1016/S0140-6736(97)11124-2
[2] Mehl AL, Thomson V. The Colorado newborn hearing screening project, 1992-1999: on the threshold of effective population-based universal newborn hearing screening[J]. Pediatrics, 2002, 109(1): E7. doi: 10.1542/peds.109.1.e7
[3] Hilgert N, Smith RJ, Van Camp G. Function and expression pattern of nonsyndromic deafness genes[J]. Curr Mol Med, 2009, 9(5): 546-564. doi: 10.2174/156652409788488775
[4] Mahboubi H, Dwabe S, Fradkin M, et al. Genetics of hearing loss: where are we standing now?[J]. Eur Arch Otorhinolaryngol, 2012, 269(7): 1733-1745. doi: 10.1007/s00405-011-1910-6
[5] Cohen M, Phillips JA 3rd. Genetic approach to evaluation of hearing loss[J]. Otolaryngol Clin North Am, 2012, 45(1): 25-39. doi: 10.1016/j.otc.2011.08.015
[6] Korver AM, Admiraal RJ, Kant SG, et al. Causes of permanent childhood hearing impairment[J]. Laryngoscope, 2011, 121(2): 409-416. doi: 10.1002/lary.21377
[7] 韩冰, 李倩, 纵亮, 等. 新生儿听力及基因联合筛查临床实践及筛查模式研究[J]. 中华耳科学杂志, 2013, 11(3): 380-383. https://www.cnki.com.cn/Article/CJFDTOTAL-ZHER201303013.htm
[8] Wang Q, Xiang J, Sun J, et al. Nationwide population genetic screening improves outcomes of newborn screening for hearing loss in China[J]. Genet Med, 2019, 21(10): 2231-2238. doi: 10.1038/s41436-019-0481-6
[9] 阮宇, 文铖, 赵雪雷, 等. 75649例新生儿耳聋基因筛查及确诊者随访结果分析[J]. 中华耳科学杂志, 2019, 17(5): 661-669. https://www.cnki.com.cn/Article/CJFDTOTAL-ZHER201905010.htm
[10] Dai P, Huang LH, Wang GJ, et al. Concurrent Hearing and Genetic Screening of 180, 469 Neonates with Follow-up in Beijing, China[J]. Am J Hum Genet, 2019, 105(4): 803-812. doi: 10.1016/j.ajhg.2019.09.003
[11] Wen C, Yang X, Cheng X, et al. Optimized concurrent hearing and genetic screening in Beijing, China: A cross-sectional study[J]. Biosci Trends, 2023, 17(2): 148-159. doi: 10.5582/bst.2023.01051
[12] Zhang J, Wang H, Yan C, et al. The Frequency of Common Deafness-Associated Variants Among 3, 555, 336 Newborns in China and 141, 456 Individuals Across Seven Populations Worldwide[J]. Ear Hear, 2023, 44(1): 232-241. doi: 10.1097/AUD.0000000000001274
[13] 中国耳聋基因筛查与诊断临床多中心研究协作组, 全国防聋治聋技术指导组. 遗传性耳聋基因筛查规范[J]. 中华医学杂志, 2021, 101(2): 97-102.
[14] Wu CC, Tsai CH, Hung CC, et al. Newborn genetic screening for hearing impairment: a population-based longitudinal study[J]. Genet Med, 2017, 19(1): 6-12. doi: 10.1038/gim.2016.66
[15] 黄卫彤, 朱茂灵, 覃卫娟, 等. 广西壮族自治区南宁市新生儿 GJB2 致聋基因的携带研究[J]. 现代检验医学杂志, 2020, 35(1): 13-15, 24. https://www.cnki.com.cn/Article/CJFDTOTAL-SXYN202001004.htm
[16] 刘清明, 田野, 於娟娟, 等. 新生儿耳聋基因筛查阳性患儿随访研究[J]. 中华耳鼻咽喉头颈外科杂志, 2019, 54(12): 881-887.
[17] 文铖, 黄丽辉, 解舒婷, 等. 中国部分地区新生儿耳聋基因筛查现况调查[J]. 临床耳鼻咽喉头颈外科杂志, 2020, 34(11): 972-977. https://lceh.whuhzzs.com/article/doi/10.13201/j.issn.2096-7993.2020.11.003
[18] Li L, Lu J, Tao Z, et al. The p. V37I exclusive genotype of GJB2: a genetic risk-indicator of postnatal permanent childhood hearing impairment[J]. PLoS One, 2012, 7(5): e36621. doi: 10.1371/journal.pone.0036621
[19] 孙毅, 刘雅琳, 刘晓莉, 等. 山东省9147例新生儿耳聋基因和听力联合筛查结果分析[J]. 听力学及言语疾病杂志, 2020, 28(5): 510-514. https://www.cnki.com.cn/Article/CJFDTOTAL-TLXJ202005006.htm
[20] 吴海燕, 鲍志宇, 杨静静, 等. 济宁地区523006例新生儿耳聋基因筛查的分析[J]. 中华耳科学杂志, 2022, 20(1): 67-71. https://www.cnki.com.cn/Article/CJFDTOTAL-ZHER202201013.htm
-
| 引用本文: | 阮宇, 程晓华, 张伟, 等. 23项新生儿耳聋基因筛查突变频谱分析[J]. 临床耳鼻咽喉头颈外科杂志, 2024, 38(4): 267-272. doi: 10.13201/j.issn.2096-7993.2024.04.001 |
| Citation: | RUAN Yu, CHENG Xiaohua, ZHANG Wei, et al. Mutation spectrum analysis of 23-site chip neonatal deafness genetic screening[J]. J Clin Otorhinolaryngol Head Neck Surg, 2024, 38(4): 267-272. doi: 10.13201/j.issn.2096-7993.2024.04.001 |
- Figure 1.
- Figure 2.



 下载:
下载: